Cores

Administration
Co-Leaders: Kirsten Moysich, PhD, MS; Kunle Odunsi, MD, PhD
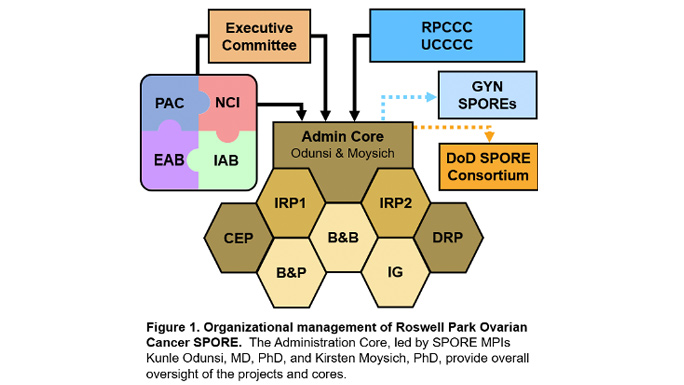
The overall objective of the Roswell Park-University of Chicago Ovarian Cancer SPORE Administration Core is to provide leadership, direction, scientific and fiscal oversight, and administrative services to enhance research productivity and maintain a stimulating research environment conducive to translational research in ovarian cancer. The Administration Core will catalyze, oversee, and support interactions amongst SPORE components, external and internal collaborators, other SPOREs, patient advocates, and the National Cancer Institute.
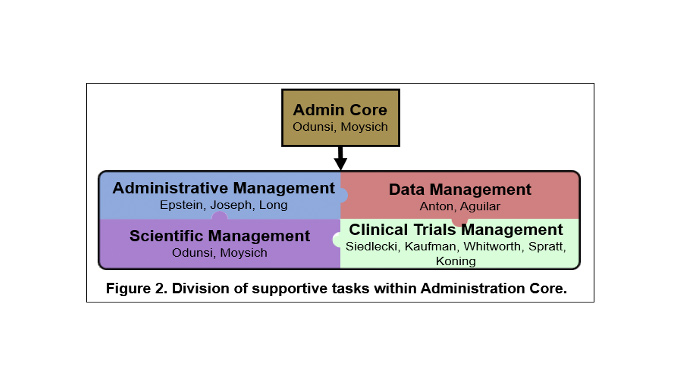
In order to achieve the overall objective of managing SPORE resources and maintaining high quality in all components, the Administration Core will be organized into four main operational areas: 1) Leadership; 2) Administrative Management; 3) Clinical Trials, Data, and Communications Management; and 4) Scientific Management. Management and support are accomplished through an organizational framework that takes advantage of highly experienced leadership that is integrated within Roswell Park Comprehensive Cancer Center and the University of Chicago Comprehensive Cancer Center, an oversight executive committee for planning and evaluation of activities, and scientific and patient advocate advisors. The functions performed by the Administration Core will provide greater integration and efficiency across the program and will result in lower operating costs for each individual SPORE component.
Biospecimen and Pathology
Co-Leaders: Carl Morrison, DVM, MD; Mark Lingen, DDS, PhD
The Roswell Park-University of Chicago Ovarian Cancer SPORE Biospecimen & Pathology Core (BPC) combines a centralized mechanism for expert tissue handling services and performance of tissue-based assays by highly qualified, experienced personnel with a state-of-the-art integrated informatics infrastructure which provides tools for tracking biospecimen collection, histopathological evaluation, and clinical annotation of samples with linkage to immunogenomic and other translational data generated by the Individual Research Projects proposed in this renewal application.
The Core will collect, de-identify, process, bank, and distribute annotated, high-quality tumor and blood samples from patients participating in SPORE-sponsored clinical studies. In addition, the BPC will provide investigators with specialized technical and interpretative histopathology services including expertise in multiplexed immunohistochemistry (IHC) and imaging mass cytometry (IMC), tissue microarray construction, laser capture microdissection, digital imaging, and quantitative pathology review of slides, tumor-infiltrating lymphocytes assessment, tissue QC/QA and preparation for next-gen sequencing-based assays (whole exome seq, RNAseq, TCRseq, MethylSEQ), and oversight on optimal utilization of each biospecimen. The Core will also be instrumental in testing patient biospecimens to determine eligibility for clinical trials and directly add to the bi-directional translational science by a detailed immunological annotation of ovarian tumors.
Biostatistics and Bioinformatics
Co-Leaders: Alan Hutson, PhD; Song Liu, PhD; Theodore Karrison, PhD
The Biostatistics and Bioinformatics Core of the Roswell Park-University of Chicago Ovarian Cancer SPORE will assist basic, translational and clinical researchers of the SPORE with proper formulation, refinement and execution of study objectives by applying the appropriate biostatistics and bioinformatics analyses, and providing the appropriate interpretation of their results, in terms of both strengths and limitations. We will ensure a priori that Developmental Research Program and Career Enhancement Program projects are feasible from a biostatistical and bioinformatics perspective.
The Biostatistics and Bioinformatics Core will ensure that appropriate data are collected; the data are of the highest quality and meet regulatory standards for privacy; and the data are those required to answer a study question, necessary for administrative reporting, needed to establish protocol compliance, and/or will be included in a manuscript. We will prepare Data Safety Monitoring Board reports and post clinical trial results to clinicaltrials.gov in a timely and compliant manner and ensure compliance to NIH policy and assist with data sharing of SPORE data via the NCI Cancer Research Data Commons infrastructure and similar NIH-designated data repositories.
Immunogenomics
Co-Leaders: Per Basse, MD, PhD, DM.Sci.; Junko Matsuzaki, PhD
The objective of the Immunogenomics Core is to perform comprehensive immunophenotyping, functional analyses, and genetic and epigenetic monitoring in peripheral blood (PB) and the tumor microenvironment (TME) and investigate tumor antigen-specific immune responses for the immunotherapeutic approaches being developed and tested in Individual Research Projects (IRP) 1 and 2 of the Roswell Park-University of Chicago Ovarian Cancer SPORE. It will be also responsible for immunologic and genomic assays for Career Enhancement Program (CEP) and Developmental Research Program (DRP). The specific aims of the Immunogenomics Core are to:
- Design and optimize immunologic and genomic assays for IRPs, CEP and DRP, and provide state-of-the-art immunophenotyping, transcriptome analysis, TCR sequencing, epigenetic changes and functional analyses of immune cells in PB and TME treated with immunotherapy.
- Run whole-exome sequencing for tumor and PB samples to identify mutations and HLA types and confirm T-cell reactivity against the predicted neoantigens for DC-peptide vaccine in IRP2.
The Immunogenomics Core will develop new methodologies/markers and make them available to SPORE investigators. It also has a significant educational role, working with all SPORE PIs, technicians and young investigators on assay development, proper use of instrumentation, and interpretation of their data.
