Mouse brain imaged from the microscopic to the macroscopic level
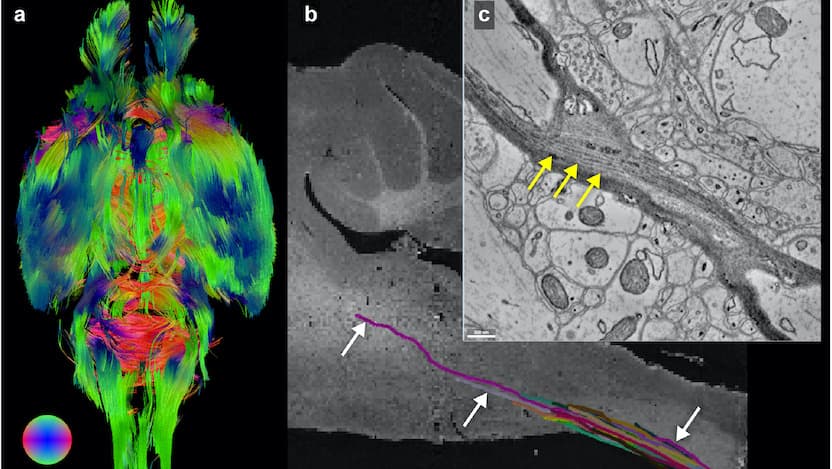
Researchers at the University of Chicago and the U.S. Department of Energy’s (DOE) Argonne National Laboratory have leveraged existing advanced X-ray microscopy techniques to bridge the gap between MRI (magnetic resonance imaging) and electron microscopy imaging, providing a viable pipeline for multiscale whole brain imaging within the same brain. The proof-of-concept demonstration involved imaging an entire mouse brain across five orders of magnitude of resolution, a step which researchers say will better connect existing imaging approaches and uncover new details about the structure of the brain.
The advance, which was published on June 9 in NeuroImage, will allow scientists to connect biomarkers at the microscopic and macroscopic level, improving the resolution of MRI imaging and providing greater context for electron microscopy.
“Our lab is really interested in mapping brains at multiple scales to get an unbiased description of what brains look like,” said senior author Narayanan “Bobby” Kasthuri, MD, Assistant Professor of Neurobiology at UChicago and neuroscience researcher at Argonne. “When I joined the faculty here, one of the first things I learned was that Argonne had this extremely powerful X-ray microscope, and it hadn’t been used for brain mapping yet, so we decided to try it out.”
The microscope uses a type of imaging called synchrotron-based X-ray tomography, which can be likened to a “micro-CT”, or micro-computerized tomography scan. Thanks to the powerful X-rays produced by the synchrotron particle accelerator at Argonne, the researchers were able to image the entire mouse brain — roughly one cubic centimeter — at the resolution of a micron, 1/10,000 of a centimeter. It took roughly six hours to collect images of the entire brain, adding up to around 2 terabytes (TB) of data. This is one of the fastest approaches for whole brain imaging at this level of resolution.
MRI can quickly image the whole brain to trace neuronal tracts, but the resolution isn’t sufficient to observe individual neurons or their connections. On the other end of the scale, electron microscopy (EM) can reveal the details of individual synapses, but generates an enormous amount of data, making it computationally challenging to look at pieces of brain tissue larger than a few micrometers in volume. Existing techniques for studying neuroanatomy at the micrometer resolution typically are either merely two-dimensional or use protocols that are incompatible with MRI or EM imaging, making it impossible to use the same brain tissue for imaging at all scales.
The researchers quickly realized that their new micro-CT, or μCT, approach could help bridge this existing resolution gap. “There have been a lot of imaging studies where people use MRI to look at the whole brain level and then try to validate those results using EM, but there’s a discontinuity in the resolutions,” said first author Sean Foxley, PhD, Research Assistant Professor at UChicago. “It’s hard to say anything about the large volume of tissue you see with an MRI when you’re looking at an EM dataset, and the X-ray can bridge that gap. Now we finally have something that can let us look across all levels of resolution seamlessly.”
Combining their expertise in MRI and EM, Foxley, Kasthuri, and the rest of their team opted to attempt mapping a single mouse brain using these three approaches. “Why did we choose the mouse brain? Because it fits in the microscope,” Kasthuri said with a laugh. “But also, the mouse is the workhorse of neuroscience; they’re very useful for analyzing different experimental conditions in the brain.”
After collecting and preserving the tissue, the team placed the sample in an MRI scanner to collect structural images of the entire brain. Next, it was placed on a rotating stage in the μCT scanner at the Advanced Photon Source, a DOE Office of Science User Facility, to collect the CT data before specific regions of interest were identified in the brainstem and cerebellum for targeting for EM.
After months of data processing and image tracing, the researchers determined that they were able to use the structural markers identified on the MRI to localize specific neuronal subgroups in designated brain regions, and that they could trace the size and shape of individual cell bodies. They could also trace the axons of individual neurons as they traveled through the brain, and could connect the information from the μCT images with what they saw at the synaptic level with the EM.
This approach, the team says, will not only be helpful for imaging the brain at the μCT resolution, but also for informing MRI and EM imaging.
“Imaging a 1-millimeter cube of the brain with EM, which is the equivalent to about the minimum resolution of an MRI image, produces almost a million gigabytes of data,” Kasthuri said. “And that’s just looking at a 1-millimeter cube! I don’t know what’s happening in the next cube, or the next, so I don’t really have context for what I’m seeing with EM. MRI can provide some context except that scale is too big to bridge. Now this μCT gives us that needed context for our EM work.”
On the other end of the scale, Foxley is excited about how this approach can be helpful for understanding the living brain through MRI. “This technique gives us a really clear way to identify changes in the microstructure of the brain when there is a disease or injury present,” he said. “So now we can start looking for biomarkers with the μCT that we can then trace back to what we see on the MRI in the living brain. The X-ray lets us look at things on the cellular level, so then we can ask, what changed at the cellular level that produced a global change in the MRI signal on a macroscopic level?”
The researchers are already using this technique to begin exploring important questions in neuroscience, looking at the brains of mice that have been genetically engineered to develop Alzheimer’s disease to see if they can trace the Ab plaques seen with μCT back to measurable changes in MRI scans, especially in early stages of the disease.
Importantly, because this work was done at the national laboratory, this resource will be open and freely accessible to other scientists around the world, making it possible for researchers to begin asking and answering questions that span the whole brain and reach down to the synaptic level.
At the moment, however, the UChicago team is most interested in continuing to refine the technique. “The next step is to do an entire primate brain,” said Kasthuri. “The mouse brain is possible, and useful for pathological models. But what I really want to do is get an entire primate brain imaged down to the level of every neuron and every synaptic connection. And once we do that, I want to do an entire human brain.”
The study, “Multi-modal imaging of a single mouse brain over five orders of magnitude of resolution,” was supported by a Technical Innovation Award from the McKnight Foundation, a Brain Initiative grant (U01 MH109100), the National Science Foundation, the National Institute of Neurological Disorders and Stroke (F31NS113571), and the National Institutes of Health (R01EB026300, S10-OD025081, S10-RR021039, and P30- CA14599). Additional authors include Vandana Sampathkumar, Scott Trinkle, Anastasia Sorokina, Katrina Norwood, and Patrick La Riviere of UChicago and Vincent De Andrade of Argonne National Laboratory.
